


| People | Alumni | Positions | Events |
| Current Projects |
| Publications |
| Theses |
| Computer Facilities |
 |
Home |
|
 |
Imprint |  |
Sitemap |  |
Intranet |  |
Contact |  |
Links |  |
 |
 |
© 2010 - 2026 Thallinger Lab · Institute of Biomedical Informatics ·
Graz University of Technology · Stremayrgasse 16/I, 8010 Graz, Austria Tel +43-316-873-5343 · Fax +43-316-873-105343 · URL http://genome.tugraz.at · |
| GENERAL DESCRIPTION |
|
The accurate measurement of the lipidome permits insights into physiological and pathological processes.
Of the present high-throughput technologies, LC-MS especially bears potential of monitoring quantitative
changes in hundreds of lipids simultaneously. In order to extract valuable information from huge amount
of mass spectrometry data, the aid of automated, reliable highly sensitive and specific analysis algorithms
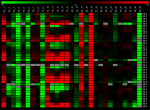
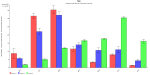
is indispensable. We present here a novel approach for the quantitation of LC-MS data. The new algorithm obtains its analytical power by two major innovations: 1) a 3D algorithm that confines the peak borders in m/z and time direction and 2) the use of the theoretical isotopic distribution of an analyte as selection/exclusion criterion. The algorithm is integrated in the Lipid Data Analyzer (LDA) application, which additionally provides standardisation, a statistics module for results analysis, a batch mode for unattended analysis of several runs and a 3D viewer for the manual verification. The statistics module offers sample grouping, tests between sample groups and export functionalities, where the results are visualised by heat maps and bar charts. The presented algorithm has been applied to data from a controlled experiment and to biological data, containing analytes distributed over an intensity range of 10^6. Our approach shows improved sensitivity and an extremely high positive predictive value compared to existing methods. Consequently, this application is a valuable improvement in the high-throughput analysis of lipids. |
CITATION |
|
1: Hartler J, Trötzmüller M, Chitraju C, Spener F, Köfeler HC, Thallinger GG. Lipid Data Analyzer: unattended identification and quantitation of lipids in LC-MS data. Bioinformatics. 2011 Feb 15, 27(4), 572-577. Epub 2010 Dec 17. PM:21169379 |

|
Support: |
|
|
| DESCRIPTION | NEWS | DOCUMENTATION | FAQ | LICENSE | DOWNLOAD |